Initialize NeuroAnalyzer
using NeuroAnalyzer
using Plots
using Wavelets
using ContinuousWavelets
eeg = load("files/eeg.hdf")
e10 = epoch(eeg, ep_len = 10)using NeuroAnalyzer
using Plots
using Wavelets
using ContinuousWavelets
eeg = load("files/eeg.hdf")
e10 = epoch(eeg, ep_len = 10)What Are Wavelets?
Wavelets are short, oscillatory waveforms that decay rapidly to zero. Unlike infinite sine waves used in Fourier analysis, wavelets are localized in both time and frequency, making them ideal for analyzing signals with transient or sharp features - such as edges in images, bursts in EEG data, or sudden changes in audio.
Core Properties
Wavelets vs. Fourier & STFT: A Comparison
| Method | Time Localization | Frequency Localization | Best Use Case |
|---|---|---|---|
| Fourier Transform | No | Global | Stationary signals (e.g., pure tones) |
| STFT | Fixed windows | Frequency bands | Quasi-stationary signals (e.g., music) |
| Wavelet Transform | Adaptive | Scale-dependent | Transient signals (e.g., edges, spikes) |
Denoising - Method: Wavelet coefficient thresholding (wavelet shrinkage). - Undesired noise is removed by adaptively thresholding wavelet coefficients corresponding to noise-dominated frequencies. - Thresholding Techniques: - Fixed-form threshold: \(\sqrt{2 \log(n)}\) (where \(n\) is the number of samples). - Adaptive methods: - Stein’s Unbiased Risk Estimate (SURE): Minimizes the risk of over- or under-thresholding. - Minimax thresholding: Optimizes for the worst-case scenario to ensure robustness.
Spectral Analysis
Wavelets excel at analyzing non-stationary signals by capturing time-localized frequency features:
Compression
Feature Extraction & Anomaly Detection
Why Wavelets?
Time-Frequency Precision: Pinpoint when and what frequencies occur in a signal.
Versatility: Used in image compression, signal denoising, biomedical analysis (EEG/ECG), and geophysical data processing.
Example: In EEG analysis, wavelets can isolate slow alpha waves (high scales) and detect high-frequency spikes (low scales) simultaneously.
Visualizing the Difference
Imagine analyzing an EEG signal:
\[ F = \frac{C_f}{s \times \mathrm{d}t} \]
where \(F\) is frequency, \(s\) is scale, and \(\mathrm{d}t\) is the sampling interval.
Scaling a wavelet means stretching or compressing its shape along the time (or space) axis by adjusting the scale parameter (sss). This directly affects two key properties:
Width:
Central Frequency: scaling inversely affects frequency. If the wavelet’s center frequency is \(F_0\) at scale \(s = 1\), then:
Example: A wavelet centered at 20 Hz at \(s = 1\) will shift to 10 Hz at \(s = 2\) and 40 Hz at \(s = 0.5\).
Why Scaling Matters
\[ F = \frac{C_f}{s \times \mathrm{d}t} \]
where \(C_f\) is the wavelet’s center frequency (a constant), and \(\mathrm{d}t\) is the sampling interval.
Practical Analogy
Think of wavelets as adjustable lenses:
mw2 = generate_morlet(100, 2, 1) # 2 Hz
mw10 = generate_morlet(100, 10, 1) # 10 Hz
t = linspace(-1, 1, length(mw2))NeuroAnalyzer.plot(t, mw2)Morlet wavelet, 2 Hz center frequency
NeuroAnalyzer.plot(t, mw10)Morlet wavelet, 10 Hz center frequency
A stretched wavelet (large \(s\)) is better suited to capturing slow, gradual changes in the signal and provides higher frequency resolution at low frequencies. A compressed wavelet (small \(s\)) captures rapid, abrupt changes and is better suited to high-frequency components where fine time resolution is needed.
Wavelet Transform: Dilations and Translations
All wavelets used in the wavelet transform are scaled (dilated/compressed) and shifted (translated) versions of a mother wavelet (\(\varphi\)).
Scaling: Achieved by dividing by the scale factor \(s\).
Large \(s\): Stretches the wavelet (low-frequency analysis). Small \(s\): Compresses the wavelet (high-frequency analysis).
Shifting: Moves the wavelet along the time/space axis to localize features.
The mother wavelet \(\varphi(t)\) is scaled and translated to analyze signals at different resolutions:
\[ \varphi(a, b) = \frac{1}{\sqrt{a}} \varphi \left ( \frac{t - b}{a} \right ) \]
Dyadic (Binary) Scaling and Translation
For efficiency and accuracy, scales (\(a\)) and translations (\(b\)) are often chosen as powers of two:
Advantages:
The number of cycles in the wavelet affects time-frequency precision:
| Cycles | Temporal Precision | Frequency Precision | Use Case |
|---|---|---|---|
| Few (2-6) | High (sharp localization) | Low (broad frequency range) | Detecting transients or edges |
| Many (10-15) | Low (smeared localization) | High (narrow frequency range) | Analyzing stationary rhythms |
Practical Guidelines
Key Insight
The choice of cycles depends on your objective:
Example:
For EEG analysis, you might use:
Here’s a clear, step-by-step guide to calculating the number of cycles and FWHM (Full Width at Half Maximum) for wavelets, with practical context:
The number of cycles (ncyc) determines how many oscillations the wavelet completes within its Gaussian envelope. You can generate a range of cycle numbers using either linear or logarithmic spacing:
Linear Spacing: Use when you want evenly distributed cycle counts across frequencies.
lowest_ncyc = 2
highest_ncyc = 10
number_of_frequencies = 5
linspace(lowest_ncyc, highest_ncyc, number_of_frequencies)5-element Vector{Float64}:
2.0
4.0
6.0
8.0
10.0Logarithmic Spacing: Use when you want exponentially increasing cycles (e.g., for better frequency resolution at higher frequencies).
lowest_ncyc = 2
highest_ncyc = 10
number_of_frequencies = 5
logspace(lowest_ncyc, highest_ncyc, number_of_frequencies)5-element Vector{Float64}:
2.0
2.990697562442441
4.47213595499958
6.68740304976422
10.0When to Use Which?
FWHM measures the frequency smoothing applied by the wavelet. It is the width of the wavelet’s frequency response at half its peak amplitude.
Steps to Calculate FWHM
Example: For a 10 Hz wavelet, FWHM ≥ 100 ms (since 1 cycle = 100 ms).
Why FWHM Matters
Practical Example
Suppose you’re analyzing a signal with frequencies from 1 Hz to 50 Hz:
logspace(1, 50, 10) to emphasize lower frequencies.Key Takeaway:
Use logarithmic spacing for cycles if you need finer resolution at lower frequencies. FWHM ensures the wavelet’s frequency smoothing is appropriate for the signal’s characteristics.
Mike X Cohen Morlet wavelet equation:
\[ mw = \exp(2 \times 1i \times \pi \times f \times t) \times \exp(\frac{-4 \times \ln(2) \times t^2}{h^2}) \]
where \(h\) is FHWM in seconds
| Wavelet | Key Characteristics |
|---|---|
| Morlet | Sine wave modulated by a Gaussian. Acts as a band-pass filter when convolved. |
| Mexican Hat | (Ricker/Marr wavelet) Second derivative of a Gaussian. Useful for edge detection. |
| Haar | Simplest wavelet; piecewise constant. Discontinuous, but computationally efficient. |
| Daubechies | Family of orthogonal wavelets with compact support. Balances smoothness and locality. |
| Coiflets | Similar to Daubechies but with vanishing moments for both wavelet and scaling functions. |
| Biorthogonal | Asymmetric wavelets for reconstruction and analysis. Used in image compression. |
| Symlets | Nearly symmetric Daubechies wavelets. Reduces phase distortion. |
Key Insight
The choice of wavelet type depends on the application - e.g., Morlet for frequency analysis, Haar for simplicity, or Daubechies for smooth, localized features.
Wavelet transforms can be implemented using Finite Impulse Response filters , enabling efficient computation of both forward (decomposition) and inverse (reconstruction) transforms. This filter-based approach is foundational in applications like signal processing and image compression.
Key Properties
Why It Matters
Flexibility: Biorthogonal wavelets can be designed to have symmetric filters, which simplify boundary handling and reduce artifacts.
Efficiency: FIR filter banks enable fast computation, making real-time processing feasible.
Practical Example:
In image compression, biorthogonal wavelets (e.g., Cohen-Daubechies-Feauveau 9/7) are widely used due to their balance of smoothness, localization, and computational efficiency.
Definition and Characteristics
The Morlet wavelet is a complex-valued wavelet formed by a sine wave (carrier) modulated by a Gaussian envelope (tapering function).
Mathematical Form:
\[ \psi(t) = e^{i \omega_0 t} \cdot e^{-t^2 / 2} \]
where:
mw = generate_morlet(100, 1, 1)
t = linspace(-1, 1, length(mw2))
NeuroAnalyzer.plot(t, mw)Morlet wavelet
Key Features
Applications:
Visualization
The Morlet wavelet resembles a damped oscillation:
Example Use Case:
In neuroscience, Morlet wavelets are often used to analyze brainwave rhythms (e.g., alpha or gamma waves) due to their ability to isolate frequency bands while preserving temporal resolution.
Overview
The Wavelet Transform decomposes a signal into time-frequency space by convolving it with scaled and shifted versions of a mother wavelet. This process yields a 3D representation (scalogram) of scale × time (t) × amplitude, revealing how frequency content evolves over time.
Key Concepts
Types of Wavelet Transforms
| Transform | Description | Applications |
|---|---|---|
| Continuous (CWT) | Uses all possible scales, producing dense time-frequency data. | Time-frequency analysis, localized filtering. |
| Discrete (DWT) | Uses discrete scales/positions (e.g., dyadic), reducing computational complexity. | Denoising, compression. |
| Stationary (SWT) | Shift-invariant version of DWT, avoids downsampling. | Signal alignment, pattern recognition. |
| Multiresolution (MRA) | Hierarchical decomposition into approximation (low-pass) and detail (high-pass) components. | Feature extraction, hierarchical analysis. |
Multiresolution Analysis (MRA)
Output: The Scalogram
Practical Considerations
Frequency Limits:
Phase Sensitivity:
Advantages & Limitations
| Pros | Cons |
|---|---|
| Time-frequency localization. | Requires band-pass filtering. |
| Adaptive resolution (MRA). | Phase information needs extra steps (e.g., Hilbert transform). |
| Efficient for non-stationary signals. | Sensitive to wavelet choice and parameters. |
Example Workflow
Discrete Wavelet Transform (DWT) decomposes a signal into approximation (low-frequency) and detail (high-frequency) components using discrete scales (\(a\)) and positions (\(b\)). This reduces computational complexity and data volume compared to CWT. Key Features
Default Wavelet: WT.haar (can be changed via the wt parameter; see Wavelets.jl for options).
Efficiency: Discrete \(a\) and \(b\) values enable faster analysis and reduced redundancy.
Multi-Level Decomposition
Methods for Choosing \(a\) and \(b\)
Redundant Wavelet Transform (Frames):
Orthonormal Bases / Multi-Resolution Analysis (MRA):
DWT Variants
| Variant | Description | Advantages | Redundancy |
|---|---|---|---|
| Standard DWT | Downsampling at each level; compact but not shift-invariant. | Computationally efficient. | Low (non-redundant). |
| Stationary WT (SWT) | Shift-invariant: No downsampling; filter coefficients upsampled by \(2^{(j-1)}\) at level \(j\). | Ideal for feature extraction/denoising. | High (N-level redundancy). |
| Autocorrelation WT (ACWT) | Redundant: No downsampling; retains all information. | Preserves signal integrity. | High. |
By default, the maximum level of decomposition is calculated automatically.
Performing 2-level stationary discrete wavelet decomposition:
dwd(e10,
ch="Fp1",
type=:sdwt,
l=2)Performing 2-level discrete autocorrelation wavelet transform (ACDWT):
dwd(e10,
ch="all",
type=:acdwt,
l=2)Since 1 + sum(2 .^ (1:l)) coefficients are produced for each channel and every epoch, resulting object may consume a lot of memory.
In the output object, coefficients are structured by rows in the following sequence:
Signal can be reconstructed from DWT coefficients using the Inverse Discrete Wavelet Transform (IDWT)
IDWT is the process of reconstructing the original signal from its DWT coefficients—the approximation (low-frequency) and detail (high-frequency) components. When no coefficients are altered, IDWT perfectly restores the original signal, making it a powerful tool for applications like denoising, compression, and feature extraction.
Reconstructing the signal using the 4th coefficients only:
dc = dwd(e10,
ch="Fp1",
type=:acdwt,
l=2)
# get data for the 5th epoch
dc = dc[1, :, :, 5]
# reconstruct the signal
s_reconstructed = idwd(dc,
type=:acdwt,
c=[4])Tip: The 1st coefficient is the original signal.
Original signal:
NeuroAnalyzer.plot(e10.epoch_time,
e10.data[1, :, 5])Reconstructed signal:
NeuroAnalyzer.plot(e10.epoch_time,
s_reconstructed)Plotting DWD coefficients:
dc = dwd(e10,
ch="Fp1",
type=:acdwt,
l=2)
# get data for the 5th epoch
dc = dc[1, :, :, 5]
plot_dwc(dc,
t=e10.epoch_time)Default wavelet for all continuous wavelet transform function is wavelet(Morlet(2π), β=2). It can be changed using the wt parameter. See ContinuousWavelets.jl documentation for the list of available wavelets.
CWT provides a continuous mapping of the signal in time and frequency.
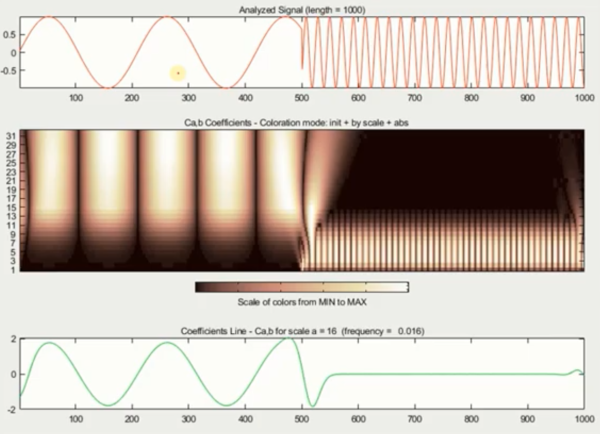
Coefficients may have redundant information since same feature may be captured by multiple scales.
CWT is suitable for detailed analysis and visualization of signals. NeuroAnalyzer uses CWT to calculate and plot scalogram.
Perform continuous wavelet decomposition (CWD):
cwd(e10, ch="all");Denoising using continuous wavelet decomposition:
denoise_cwd(e10,
ch="all",
nf=50);Tip: Denoising is performed by zeroing coefficients around the noise frequency and running inverse transform to reconstruct the signal.
Denoising using discrete wavelet decomposition:
denoise_dwd(e10,
ch="all",
smooth=:undersmooth,
l=2);l is the the number of decomposition levels.
smooth is the smoothing method used (:regular thresholds all given coefficients, whereas :undersmooth smoothing does not threshold the lowest frequency subspace node of the wavelet transform).
Tip: Default denoising function (dnt) is RelErrorShrink(SoftTH()). See WaveletsExt.jl for detailed description of available denoising functions.
Before:
NeuroAnalyzer.plot(e10,
ch="Fp2",
ep=7)plot_spectrogram(e10,
ch="Fp2",
ep=7,
flim=(0, 20))Removing artifact at epoch 7:
e10_new = artrem_cwd(e10,
ch="Fp2",
ep=7,
tseg=(0.5, 2.5),
fseg=(0.5, 7.5))Tip: Artifact position time segment must be provided within the epoch time, hence tseg=(0.5, 2.5) above.
Tip: Artifact removal is performed by zeroing coefficients around the artifact time and frequency and running inverse transform to reconstruct the signal.
After:
NeuroAnalyzer.plot(e10_new,
ch="Fp2",
ep=7)plot_spectrogram(e10_new,
ch="Fp1",
ep=7,
flim=(0, 20))